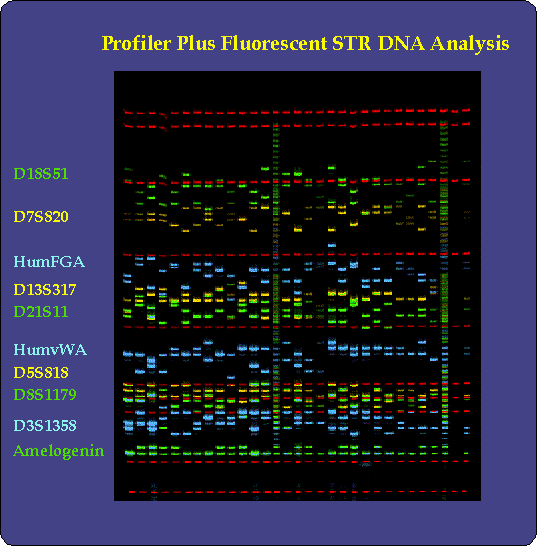
Gel electrophoretic separation of DNA fragments containing Variable Numbers of Tandem Repeats. Each vertical lane contains one DNA sample. PCR-amplified fragments have been labeled, with different probes having different fluorescent labels. The red bands are internal calibration markers. The positions of the bands reflect their different mobilities, which in turn reflect their lengths (larger fragments move more slowly in electrophoresis). Capillary electrophoresis and automatic recording are used today instead of gel electrophoresis.
DNA profiling resulted from the convergence of scientific curiosity, technological advances, and the desire to provide a social benefit. It has made a solid contribution to forensic science. However, as originally carried out, its sensitivity was limited because it required relatively large amounts of material. This restricted its application in many criminal investigations. This limitation was effectively removed by a singular development in molecular biology, the invention of the Polymerase Chain Reaction (PCR). While the original DNA profiling methods used the DNA available in the sample, the PCR massively increases the amount of DNA, theoretically without limit. It uses the molecular machinery, the enzymes, that make an exact copy of DNA during each round of cellular proliferation.
The Idiosyncratic Dr. Mullis
The “Polymerase Chain Reaction”, PCR, was invented by Dr. Kary Mullis and colleagues at the Cetus Corporation in 1983. It quickly became so important, not only in DNA profiling, but in all of molecular genetics, that Mullis won a Nobel Prize in 1993. Although invention of the PCR method is worthy of a Nobel Prize, readers of this site will know that I do not hold Mullis in unalloyed admiration. For example, he used the prestige of the Nobel Prize to undermine the link between HIV and AIDS, and to disparage the scientists who supported it. This had consequences in South Africa (described here).
Many deaths have been attributed to the refusal of South African authorities to treat HIV-positive victims with appropriate drugs because they believed Mullis, and a few other contrarians who disputed the HIV-AIDS link. Mullis also claimed not to believe in the “ozone hole” in the atmosphere, which mainstream scientists attributed to the use of certain industrial halocarbons. He was undoubtedly wrong about this too: the discontinuation of the use of these chemicals has led to the gradual shrinking of the ozone hole. It was also his opinion that climate scientists were wrong about global warming. He asserted that he believed in astrology, although that was in a somewhat humorous vein in a 1998 autobiography. Despite these counterfactual positions, he was indeed the first person to carry out (although not to envisage the principle of) the Polymerase Chain Reaction, and none of his contrarian idiosyncrasies diminishes that contribution. Mullis died in 2019 at age 74.
Using PCR to Increase the Sensitivity of DNA Profiling
The power of PCR to enhance DNA profiling can be understood using the example of the previous post, in which two individuals have different numbers of Variable Number Tandem Repeats at a particular locus of their genomic DNA. In that example, one person has 5 copies of the 5-nucleotide repeat GCTTA in one DNA strand and its complement in the other. The second person has 7 repeats.
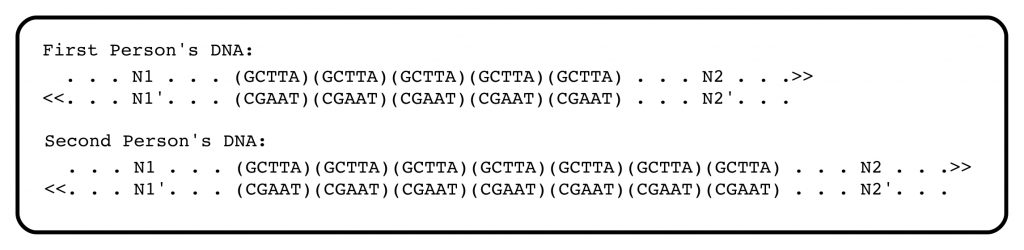

The analysis of the first person’s DNA using PCR begins by heating the DNA to cause the strands to separate — this is “heat denaturation”. A short chemically-synthesized fragment of DNA, called a primer, is added. The primer, perhaps 15 nucleotides long, is complementary to N2 (in other words, it has the same sequence as N2′). The primer binds to its complementary N2 sequence in the top strand. A DNA polymerase enzyme, which can copy DNA into its complement, is added, together with its substrates, the four deoxynucleotides. The result is the synthesis of the complement to the top strand of the DNA, beginning with the primer N2′ (in blue) and ending somewhere off to the left (it doesn’t matter where).
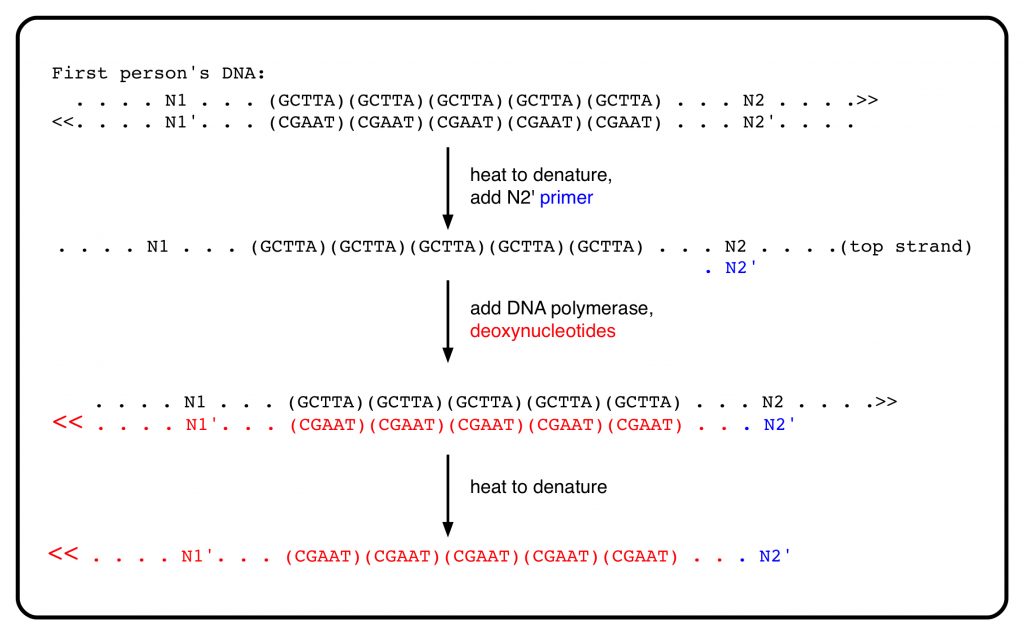
The DNA polymerase enzyme has an important, limiting, feature: It cannot initiate a new strand. It needs something to add on to, and in the PCR technique, that is provided by the experimenter in the form of the short primer. The chemical synthesis of DNA primers was another Nobel-worthy achievement of the 1960s. The need for a primer to start copying is a feature, not a failure of the system in vivo; it is a mechanism for controlling DNA replication in cells. In PCR, it allows for controlling which part of DNA will be copied by the DNA polymerase. The product of copying begins precisely with the N2′ (primer) sequence. It extends to the left of N1, but that doesn’t matter.
That’s one cycle of copying. In the next cycle, the product of the first step is denatured by heat and copied. This copying too is initiated with a synthetic primer (also in blue) that is complementary to N1′ (in other words, it has the short sequence around N1). The product of this copying ends precisely at N2, since the strand it is copying physically ends there. Now there’s a product the exact length of the distance between N1 and N2. (The intermediate product from the first cycle, which extends to the left of N1 and has an indeterminate length, is diluted out by the repeated cycles of the PCR.)
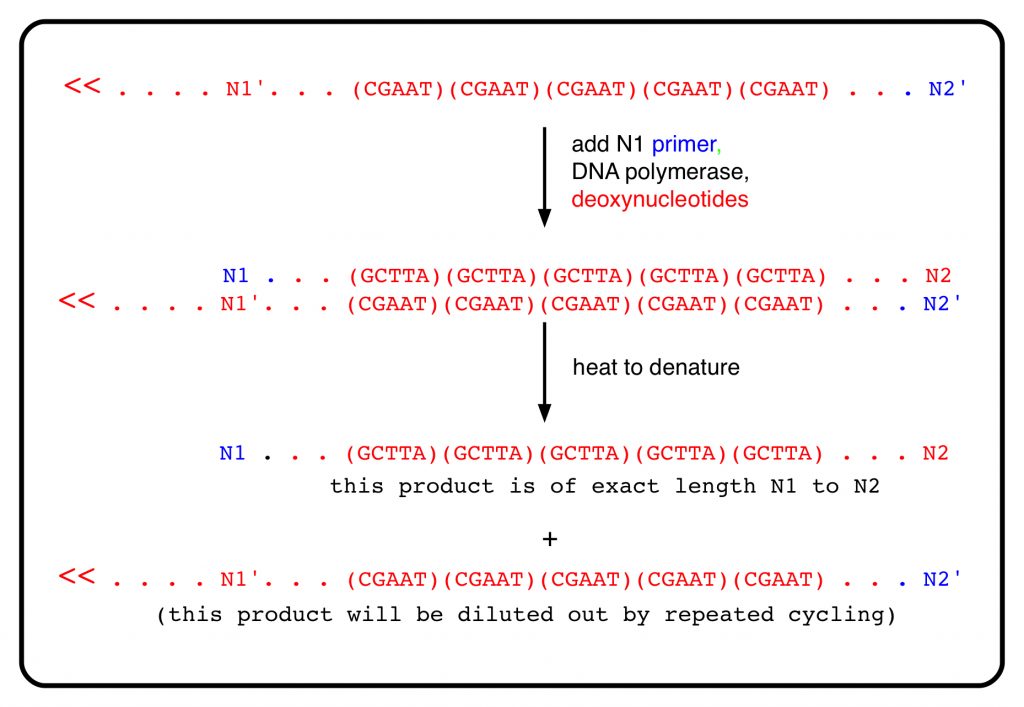

The bottom strand of DNA is copied in the same fashion. Each round of copying can theoretically double the amount of product of length equal to the distance from N1 to N2. For the first person in the comparison, the length of the product reflects its original content of 5 copies of the VNTR; for the second person, it will be 10 nucleotides longer because their genomic DNA has 7 copies of the VNTR. High-resolution, automated electrophoretic separation of the products is used to determine the VNTR pattern of the original DNA. This is the heart of the PCR-based DNA profiling method. In the picture at the top of the post, you can see differences in the distances moved of the blue fragments between lanes, reflecting differences in the number of VNTR repeats in that part of the genome for the 2 people being compared.
In forensic DNA profiling, a number of probes, each corresponding to a different part of the human genome, are used. These probes (between 13 and 20 in number, depending on the country) are chosen because they show a high level of variation within the population (‘allelic variation’). Also, the probes correspond to various chromosomes, to broadly reflect the genome. If two people happen to have the same versions of one of the VNTR fragments, as can happen, they nevertheless will differ in most of the others, providing powerful statistical evidence to distinguish them.
Effects of DNA Profiling on Justice
Because of its proven credibility and discriminating power, DNA profiling has become a standard of comparison for other types of forensic evidence. A study by the National Academy of Sciences of the USA in February 2009 provides a sobering evaluation of the reliability of various standard forms of evidence. Blood tests, bite marks, footprints, soil samples, fiber analysis and conventional fingerprints provided sufficient evidence to obtain convictions in 137 of the cases that were overturned by unequivocal DNA profiling evidence (1). It was estimated that eyewitness misidentification was a factor in over 70% of the overturned wrongful convictions. And, as in the initial criminal case solved by Jeffreys, a confession can be false.
The Innocence Project was started in 1992 at the Cardozo School of Law in New York. Its purpose is to try to get justice for the wrongly convicted. Today it is one of a number of international organizations dedicated to this purpose. Some of the cases the Innocence Project has overturned have depended on ‘DNA exonerations’. As of September 2019, the Innocence Project has obtained 365 exonerations based on DNA profiling evidence, 20 of these being death penalty cases. Although the evidence unequivocally showed these individuals to be innocent, 25% had confessed to the crimes. DNA profiling usually comes into play only in the most serious crimes, such as rape and murder, which lead to long prison sentences, or, in the United States, death. These innocent individuals had spent an average of 14 years in prison, some more than 25.
Specific cases provide a frightening view of how the criminal justice system can fail, and the ability of DNA profiling to rectify an injustice. In Canada, a 19 year-old man named David Milgaard was found guilty of raping and murdering a young nurse on the last day of January 1969. The crime took place on a miserably cold night in a back lane in the prairie city of Saskatoon. Milgaard was given a life sentence, and because he refused to admit he’d done the crime, he was repeatedly denied parole. During his 23 years in prison his mother campaigned tirelessly to have the conviction overturned, on the basis of exculpatory evidence that had not been considered at, or that had emerged since, his trial. She was unsuccessful, until she somehow gained the ear of Prime Minister Brian Mulroney, who instructed his Minister of Justice to have the Canadian Supreme Court re-evaluate the case. This resulted in Milgaard’s release from prison, although not his exoneration (the Court basically said that he shouldn’t be in prison any longer, but did not find him innocent).
What finally lifted the stain of the guilty verdict from David Milgaard’s shoulders was DNA evidence, which had been maintained in an evidence room as a dried semen stain on a pair of panties from 1969 until it was analyzed in 1997. As a bonus, the DNA evidence led to the conviction of another man, a known sex offender named Larry Fisher (2). (In retrospect, even conventional evidentiary clues should have included him in the list of suspects, and made David Milgaard a somewhat unlikely one.) DNA profiling has injected a new level of proof into criminal law that is forcing the justice system to scrutinize its methods with a critical eye (1).
When the Best Forensic Tool Fails
There are limits to any forensic tool, and DNA profiling is no exception. Evidence must still be handled properly, for example. And because the method is so sensitive, contamination is always a danger. Such limitations were invoked by the defense in asking the jury to reject DNA evidence as proof in the notorious case of O. J. Simpson. They did.
There can be other reasons for failure of PCR-based DNA profiling. In a spectacular burglary in Berlin in 2009, jewelry worth over six million euros taken from the KaDeWe department store by three masked men (3). One of the thieves left behind a latex glove from which enough material was recovered to allow DNA profiling. The results pointed to a person already in the criminal DNA database. Unfortunately for the police and fortunately for him, he has an identical twin, and no available DNA test could resolve which of the two men had worn the glove. (A total DNA sequencing approach might have done that, but could not be used because of the time, expense, and uncertainty involved.) And the thieves had been careful — no conventional fingerprints, which would have identified the brother involved, were left behind (interestingly, identical twins have different conventional fingerprints). As of 2018, no guilty verdict has been issued.
The New York Times recently carried a story about a man in Reno, Nevada, who had DNA in his body different from what he was born with (4). How did that happen? He had received a lifesaving bone marrow transplant, from a donor in Germany he never met, as a treatment for a blood cancer. The transplant was successful, and the bone marrow recipient is in remission. A colleague who works in a sheriff’s office wanted to see what effect the new bone marrow would have on his body’s DNA. Not surprisingly, the recipient’s blood had been repopulated with cells of the donor, which was the point of the transplant. But so had the cells of his semen, which are largely white blood cells! (He had had a vasectomy, so sperm cells were not present; they would have contained his own DNA.) Cells collected from his lip, cheek and tongue also contained the donor’s DNA, to variable percentages.
This result was reminiscent of a criminal case in Alaska, where semen samples indicated that the perpetrator of a rape was in the DNA database. The problem was, he was in jail at the time of the crime. The explanation was that he had earlier received a bone marrow transplant from his (non-identical-twin brother). The (non-incarcerated) brother was subsequently identified as the perpetrator of the crime.
These outcomes are the result of chimerism: DNA from stem cells in the bone marrow transplant has taken over part of the recipient’s body. As it happens, no changes in appearance or behavior have been noticed. You are still you, although some of your cells have come from someone else.
Given the almost universal use of PCR around the world, it shouldn’t be surprising that there have been some failures. Its application isn’t always well monitored. Technical incompetence is one source of concern: if the samples aren’t handled properly, or the judgment of results is faulty, there can be errors that send people to prison or worse. The element of judgment is particularly concerning because of a push to use “Artificial Intelligence” to make them, in the interest of saving money. The computer software used to make the judgment calls by AI is often a black box, and its accuracy can be difficult to determine.
A whistle-blower alerted a radio station in Houston, Texas, about bad practice in a local DNA lab, and investigation showed that technicians were routinely misinterpreting even the most basic samples (5). A young man was sentenced to 25 years for a rape that was based on incompetently and incorrectly profiled DNA (the evidence for error became clear on investigation). He was found guilty despite having a solid alibi, and despite the victim identifying quite a different body type for her attacker. No technique will withstand willful or accidental malpractice.
The Need For Continuous Improvement
In addition to dealing with the problems of misuse and misadventure outlined already, scientists are always on the lookout for useful tricks and tweaks that improve PCR-based DNA profiling. One involves the kind of DNA polymerase used. Most DNA polymerases, like other enzymes, are sensitive to heat, and are destroyed during the heating phase after every round of PCR copying. In the original method, fresh DNA polymerase enzyme had to be added for each cycle. But biologists had discovered some remarkable microorganisms that live at very high temperatures in the hot geysers of Yellowstone Park. Not surprisingly, these organisms have heat-stable enzymes, including a heat-resistant DNA polymerase. This “Taq polymerase”, named for its microbial cell of origin, Thermus aquaticus, can withstand many rounds of heating of the PCR sample, and is now used routinely.

Go To Contents
Sources cited
- Strengthening Forensic Science in the United States: A Path Forward. The National Academies Press, 2009, Washington, D. C.
- Perreaux, L. Fisher gets life with no parole for 10 years. In Saskatoon Star Phoenix, January 5, 2000, Saskatoon, SA.
- Diehl, J. 2009. Perfect genes for a robbery. In Der Spiegel, February 18, 2009.
- Murphy, H. 2019. When a DNA Test Says You’re a Younger Man, Who Lives 5,000 Miles Away. New York Times.
- Shaer, M. 2016. TheFalse Promise of DNA Testing. The Atlantic June 20165.